Introduction
Epilepsy is one of the most common neurological disorders with a prevalence of about 1% worldwide. The most frequent form of human epilepsy is mesial temporal lobe epilepsy (mTLE) (), with seizures often originating in the hippocampus as a result of abnormal electrical activity (). Brain insults that lead to aberrant neuronal wiring, for example in the hippocampus or entorhinal cortex, may cause mTLE, albeit it is still unclear in which cases insults induce epileptogenesis—and when this then clinically manifests as mTLE (, ). mTLE is commonly associated with hippocampal sclerosis, characterized by glial scarring, the loss of pyramidal neurons in the first and third regions of the Cornu ammonis (CA1 and CA3), and granule cell dispersion (GCD) within the dentate gyrus (DG) (; ). Unfortunately, it can be resistant to pharmacological treatment, thereby requiring surgery to remove epileptic lesions (), although this course of action does not always ensure lifelong freedom from seizures ().
Neuroimaging plays an important role in diagnosing and understanding epilepsy. For example, determining epilepsy biomarkers for in vivo magnetic resonance imaging (MRI) is an active area of research (), where validation with post mortem cellular-resolution imaging is critical (). Conventional histology with well-established immunohistochemistry is the gold standard for the histopathological investigation of mTLE (). Detailed information about cellular changes associated with mTLE can be drawn from cell-type-specific immunohistochemistry. In experimental models, this is important as the loss of neurons and the subsequent activation of astrocytes as well as microglia show distinct spatiotemporal patterns during disease progression, indicating specific roles in remodeling of the epileptic hippocampal network. Moreover, in mTLE patients hippocampal sclerosis comes in different flavors: neuronal loss can be partial or nonexistent, while gliosis is present. In fact, hippocampal sclerosis with only gliosis has a particularly bad postsurgical prognosis (), which renders detailed histological analysis of resected hippocampi in mTLE important for patients and their families. While these aspects highlight the biological relevance of immunohistochemical analysis in mTLE, the present study leverages immunohistochemistry to demonstrate the strengths and limitations of the microtomography’s electron-density-based contrast, to visualize individual cellular elements. However, this approach has limited out-of-plane resolution and suffers from artifacts during staining and sectioning (). As a result, any correlation with MRI can involve complex slice-to-volume registration (). Visualizing the three-dimensional arrangement of hippocampal cells is important for understanding the progression of mTLE (; ; ), but MRI is suboptimal for this task due to its relatively low spatial resolution. Even magnetic resonance microscopy lacks spatial resolution, with post mortem resolution of the order of tens of μm down to about 10 μm in special cases (; ; ). Therefore, a label- and slicing-free volumetric imaging approach with micrometer resolution is highly desirable, in that compatibility with both MRI and conventional histology workflows would allow for correlation and validation.
Hard X-ray micro- and nanotomography are attractive options for this role, as X-ray imaging can bypass sectioning or tissue clearing and provides spatial resolution below the limits of optical microscopy (; ). Soft tissues are characterized by minute differences in X-ray absorption. Therefore, for microstructure analysis beyond hard tissues such as bone, X-ray microscopy has traditionally relied on contrast agents and/or corrosion casts (; ). These approaches have significant advantages over light microscopy approaches: recent staining protocols enabled volumetric imaging and quantification of the entire vascular and tubular system of a mouse kidney, avoiding artifacts from optical distortion or morphology changes during tissue clearing (). Further, recent advances in X-ray source brilliance and phase contrast imaging have increased spatial and density resolution to allow imaging of unstained brain tissue with micrometer (; , ) or even nanometer () resolution. X-ray microtomography as an intermediary step in standard histopathological preparation has allowed for extending conventional histology to the third dimension (), and it also provides enhanced X-ray contrast (; , ). Thanks to this maturation in the use X-ray microtomography for histopathological analysis, this family of techniques is often referred to as three-dimensional (3D)X-ray histology.
The present study evaluates 3D X-ray histology with synchrotron radiation-based X-ray microtomography to track the histopathological progression of mTLE in a mouse model. A kainate (KA) mTLE model is used, whereby a stereotactic KA injection into the hippocampus induces a pathological phenotype resembling that of humans (). Specifically, injection is followed by hippocampal sclerosis including the loss of pyramidal cells and interneurons as well as GCD and gliosis (; ; ; ). This is closely linked to the emergence of epileptic discharges (, ). Label-free 3D X-ray histology with 1.6 μm-wide voxels was used to image paraffin-embedded brain hemispheres from mice sacrificed 1, 7, 14, and 21 days after KA injection, i.e., time points representing distinct phases during epileptogenesis (). The onset of hippocampal sclerosis, namely GCD and pyramidal neuron loss, was identified with 3D X-ray histology. 3D X-ray histology data were benchmarked against corresponding sections in conventional histology. Finally, the automatic segmentation of the molecular, granular, and polymorph layers of the DG with a deep convolutional neural network enabled us to three-dimensionally quantify electron density and morphology changes during epileptogenesis.
Materials and Methods
Animals and Kainate Injection
Fifteen 9- to 12-week-old male C57BL/6N wildtype mice (Charles River, Sulzfeld, Germany) were used for this experiment. The mice were housed at room temperature in a 12-h light/dark cycle and were provided with food and water ad libitum. Animal procedures were in accordance with guidelines set out in the European Community’s Council Directive of 22 September 2010 (2010/63/EU) and were approved by the regional council (35-9185.81/G-16/38, Regierungspräsidium Freiburg, Germany).
The intrahippocampal KA mouse model has been extensively studied and validated as a model for mTLE (; ; ; ). Note that hippocampal sclerosis is closely linked to the emergence of epileptic discharges (, ). The 15 mice formed five groups of three replicates. The timeline is summarized in Figure 1. On Day 0, Group 1 were injected with saline and Groups 2–5 were injected with kainate. On Day 1, Group 2 were sacrificed. On Day 7, Groups 1 and 3 were sacrificed. On Day 14, Group 4 were sacrificed. On Day 21, Group 5 were sacrificed. The injection procedure is described in the following references (; ; ). In short, anesthetized mice were sterotactically injected with 50 nL of either 0.9% saline solution (Group 1) or 20 mM kainate solution (Tocris, Bristol, UK) in 0.9% saline (Groups 2–4) into the right dorsal hippocampus. One of the mice from Group 3 died before reaching the sacrifice time point. Due to the limited available beamtime at the beamline ID19 of the European Synchrotron Radiation Facility (Grenoble, France), a total of 22 of the 28 available brain hemispheres were measured with hard X-ray tomography, see summary in Supplementary Table 1. According to our priority list, all 14 ipsilateral hemispheres were acquired to quantify hippocampal sclerosis. Eight additional contralateral hemispheres could also be measured in the allotted time. Contralateral hemispheres were mainly included to rule out histological changes such as CA1 cell death.
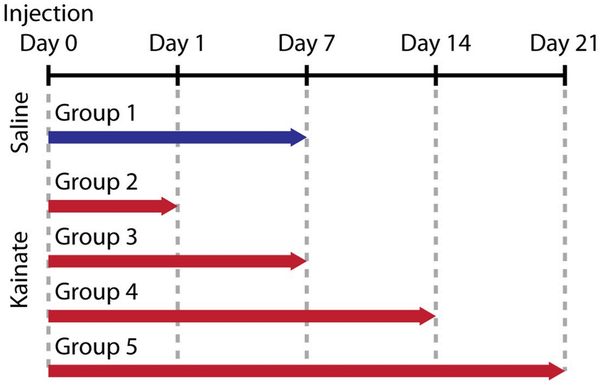
Fig. 1
Study design. Fifteen mice formed five groups of three replicates. On Day 0, Group 1 were injected with saline and Groups 2–5 were injected with kainate. On Day 1, Group 2 was sacrificed, followed by Groups 1 and 3 on Day 7, Group 4 on Day 14, and Group 5 on Day 21. The brains were extracted, fixed in paraformaldehyde, and embedded in paraffin. 3D X-ray histology was then performed prior to standard histology.
Tissue Preparation
The mice were anesthetized and transcardially perfused with 0.9% saline followed by 4% paraformaldehyde in 0.1 M phosphate buffer for 5 min, then sacrificed. Brains were extracted and postfixed in 4% paraformaldehyde with 0.1 M phosphate buffer overnight at a temperature of 4 C. After fixation, a scalpel was used to separate the brain hemispheres and remove the cerebellum and olfactory bulbs. Brain hemispheres were embedded in paraffin following the standard procedure in histopathology: the tissue was dehydrated in ascending ethanol solutions, cleared in xylene, and embedded in liquid paraffin wax (Surgipath Paraplast®, Leica Biosystems, Wetzlar, Germany) before cooling to form a solid block (). The paraffin-embedded samples were imaged with a nanotom m® (GE Sensing and Inspection Technologies GmbH, Hürth, Germany) laboratory microtomography system to identify defects such as cracks, highly absorbent debris, or trapped air. Samples with defects were melted and re-embedded in paraffin. A tissue punch with an inner diameter of 6 mm was used to extract cylindrical samples containing an entire brain hemisphere and minimal excess paraffin.
Synchrotron Radiation-based Microtomography
X-ray microtomography of the mouse brain hemispheres was performed at the European Synchrotron Radiation Facility’s (Grenoble, France) ID19 beamline. A “pink beam” () with a mean photon energy of 19 keV was used. A detector based on a 100 μm-thick Ce:LuAG scintillator, coupled via a magnifying objective to a scientific CMOS camera (pco.edge 5.5, pixel array, 6.5 μm physical pixel size) was used to achieve an effective pixel size of 1.625 μm. The resulting detector’s field of view was . A propagation distance of 280 mm was selected to allow for propagation-based phase contrast. Each brain was scanned with 4,000 projections around 360 with an exposure time of 100 ms per projection. Three or four height steps were acquired to cover the entire brain hemispheres. Including sample changes and alignment, acquisition time was about 45 min per hemisphere, i.e., 16.5 h for 22 hemispheres. Considering setup and potential beam loss or technical issues, a similar imaging experiment would require a successful application for three 8-h shifts.
Prior to reconstruction, projections were phase retrieved by using Paganin’s filter () with , whilst ring artifacts were reduced via the method described by . Tomographic reconstruction was performed in MATLAB (release 2016a, The MathWorks, Inc., Natick, MA, USA) using a filtered back-projection algorithm with the standard Ram-Lak filter (). Any overlap between height steps was determined using the Open Source registration toolbox elastix (version 4.9) (; ), and a translation transform was used with a normalized correlation coefficient as a similarity metric. Height steps were combined with linear blending within the overlapping regions.
Single-distance phase retrieval from pink beam microtomography provides semi-quantitative gray values related to the decrement of the real part of the index of refraction . For X-ray energies far from absorption edges, is proportional to the electron density . This condition is held for physically soft tissue and photon energy at around 20 keV.
A single scientist manually aligned the tomography data to the Allen Mouse Brain Common Coordinate Framework (; ) with a rigid transformation using the Open Source tools ITK-SNAP (version 3.8.0) () and transformix (version 4.9) (; ). This facilitated comparison of anatomical features across mice. Note that a rigid transformation was employed to avoid changing volume measurements. Downsampled datasets ( binned) were used to find this rigid transformation, requiring only a few minutes per hemisphere.
Volume renderings were generated in VGStudio MAX 2.1 (Volume Graphics, Heidelberg, Germany).
Immunohistochemistry and Histology
Subsequent to tomographic imaging, paraffin-embedded brain hemispheres were immersed in xylene, descending ethanol solutions, and, finally, in 4% paraformaldehyde with 0.1 M phosphate buffer. Next, 50 μm-thick coronal sections of the brain hemispheres were produced on a vibratome (VT1000S, Leica, Bensheim, Germany). Consecutive sections were stained with hematoxylin and eosin (H&E), cresyl violet (Nissl), antibodies against hexaribonucleotide binding protein-3 and ionized calcium-binding adapter molecule 1 (NeuN-Iba1), and glial fibrillary acidic protein (GFAP). For Nissl or H&E staining, sections were stained in a cresyl violet or H&E solution for 10 min and briefly rinsed with distilled water. Tissue sections were mounted with DPX medium on Superfrost glass slides (Art. No. H867.1; Carl Roth GmbH, Karlsruhe, Germany). For immunofluorescence staining, sections were treated with 0.25% TritonX-100 in 1% bovine serum albumin for 1 h, then incubated at a temperature of 4 C for 24 h with Guinea pig anti-NeuN (1:500, Synaptic Systems, Göttingen, Germany), rabbit anti-Iba-1 (1:1000, Wako Chemicals, Neuss, Germany), or rabbit anti-GFAP (1:500, Dako, Hamburg, Germany). For detection purposes, Cy2-, Cy3-, or Cy5-conjugated secondary donkey anti-guinea pig or goat anti-rabbit antibodies (1:200, Jackson ImmunoResearch Laboratories Inc., West Grove, PA, USA) were applied for three hours at room temperature, followed by rinsing in 0.1 M PB six times for 15 min. Sections were then counter-stained with DAPI (4 ,6-diamidino-2-phenylindole; 1:10,000, Roche Diagnostics GmbH, Mannheim, Germany), after which the free-floating sections were mounted on glass slides and cover-slipped with ProLong Gold (Molecular Probes, Invitrogen, Carlsbad, CA, USA).
Histological slides were scanned using an AxioImager2 microscope (Zeiss, Göttingen, Germany). A 10× objective (Plan-APOCHROMAT, Zeiss, Göttingen, Germany) and a digital camera (MR605, Zeiss, Göttingen, Germany) were used to acquire composite immunofluorescent-stained sections. These composites were then processed with the Zen (Zeiss, Göttingen, Germany) software package.
Several histology slices suffered from artifacts, including thickness variations, folds, tears, and cracks. The most artifact-free sections were selected for comparison with 3D X-ray histology. Approximate locations of the histology sections were manually found within the tomography datasets by defining a rigid transformation with three angles of rotation and three translations.
Deep Convolutional Neural Network-based Segmentation
Deep convolutional neural networks are not invariant to rotations of the input. Hence we manually rigidly aligned the tomography data to the Allen Mouse Brain Common Coordinate Framework (; ). Thanks to this coarse prealignment, augmentation of the training data can be restricted to a smaller transformation range and hence training times are reduced.
The molecular, granular, and polymorphic layers of the DG were manually segmented from the tomography data. Manual segmentations were guided by the Allen Mouse Brain Atlas (), and segmentation was performed in Amira 6.2.0 (Thermo Fisher Scientific Inc., Waltham, MA, USA). The binned tomography data were used, whilst blow (tolerance 15, Gauss width 3) and paint brush tools were employed to manually segment virtual coronal slices.
Manual labeling of an entire volumetric dataset is time-consuming and infeasible for all 22 measured brain hemispheres; for example, manually labeling a single slice takes up to three minutes, depending on the disease state. Therefore, labeling the 514 slices of a brain hemisphere dataset, i.e., at 4× binning, would require 9–26 h. Thus, a machine-learning approach was used, which was trained on a limited number of manually labeled slices from all five mouse groups. The well-known U-Net architecture was selected (). This deep convolutional neural network (DCNN) was built using PyTorch (version 1.11.0) and Python 3 (version 3.10.4), based on publicly available code ().
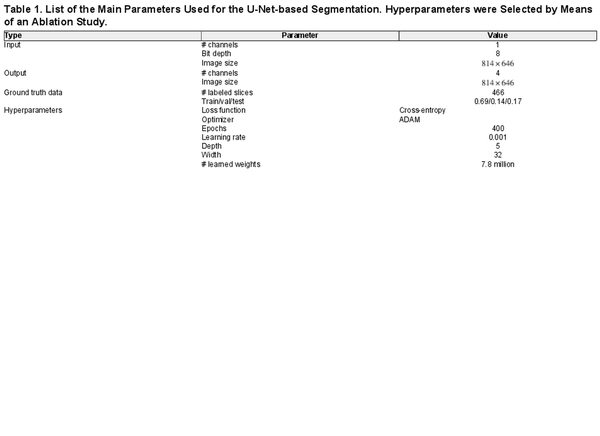
Values selected for the main U-Net-based segmentation parameters are listed in Table 1. In total, 466 manually labeled slices from 18 datasets were used. Intensity values for the slices in size, i.e. 4× binned, were reduced to 8-bit depth with the same range for all slices and hemispheres. Manually labeled slices were split into training, validation, and test at a ratio of 0.69, 0.14, and 0.17, respectively. This ratio was the result of a targeted 0.7/0.15/0.15 ratio with the constraint that at least one slice from every manually labeled brain hemisphere was present in the test set. For training, the median number of manually labeled slices per brain hemisphere was 20. The main parameters were selected based on an ablation study. The selected DCNN used a depth of 5 and width of 32, corresponding to 7.8 million learned parameters. The network was trained for 400 epochs, with cross-entropy as the loss function and ADAM as the optimizer (). Final network weights were selected from the training epoch, thus minimizing the loss function on the validation set. Training was performed at the scientific computing center sciCORE (https://scicore.unibas.ch/), University of Basel, using an NVIDIA RTX8000 GPU. Three-dimensional segmentations were produced through the slice-by-slice segmentation of virtual coronal slices.
Results
Visualizing Histopathological Changes Associated with Epilepsy
Volumetric datasets from 3D X-ray histology allowed for three-dimensional navigation within the mouse brain’s anatomy. Virtual sectioning visualized the injection site of saline or KA as well as the relevant features of the hippocampal formation, namely the Cornu ammonis and DG, see Figure 2. To enable a comparison of the healthy and epileptic brains, all measured hemispheres were coarsely aligned with the coordinate system of the Allen Mouse Brain Reference Atlas ().
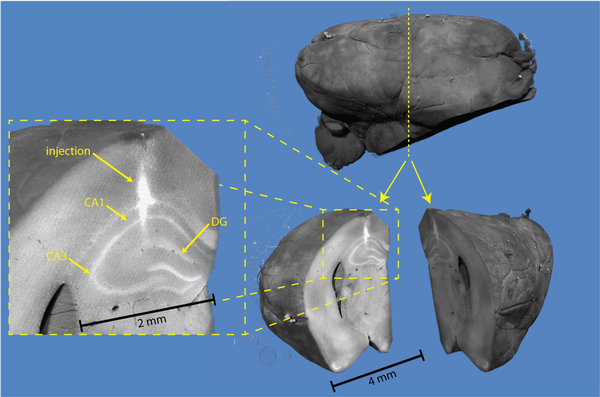
Fig. 2
3D X-ray histology of a saline-injected mouse brain hemisphere. Volume renderings show the ipsilateral hemisphere (top right) of a mouse from Group 1 sacrificed on Day 7 after saline injection. Virtual sectioning of the brain helps identify the injection site in the right dorsal hippocampus (bottom right). The pyramidal layers of the first and third fields of the Cornu ammonis (CA1 and CA3), as well as the dentate gyrus (DG) layers, are visible in the magnified view (left).
Virtual coronal slices through the 3D X-ray histology of the KA-injected brains revealed well-described histopathological changes associated with the progression of mTLE, as illustrated in Figure 3. Despite the label-free approach, hippocampal sclerosis was evidenced by GCD as well as the loss of neurons in the pyramidal layers of the CA1 and CA3 regions. The extent of hippocampal sclerosis increased over time, from the point of injection through sacrifice. In corresponding contralateral hemispheres, the pyramidal layers of the CA1 and CA3 regions were identifiable by a continuous cell layer of increased density for all mice imaged at the selected time points. The loss of pyramidal neurons in these regions was characterized by an initial increase in individual cell density (see Day 1 postinjection, right side), followed by a widening of the pyramidal layer, a decrease in density, and the loss of the layers’ contrast relative to the surrounding strata. Pyramidal neuron loss in the CA1 field was identified for mice sacrificed on Day 7 after the KA injection, while neuron loss in the CA3 field was first observed at Day 14 after the KA injection.
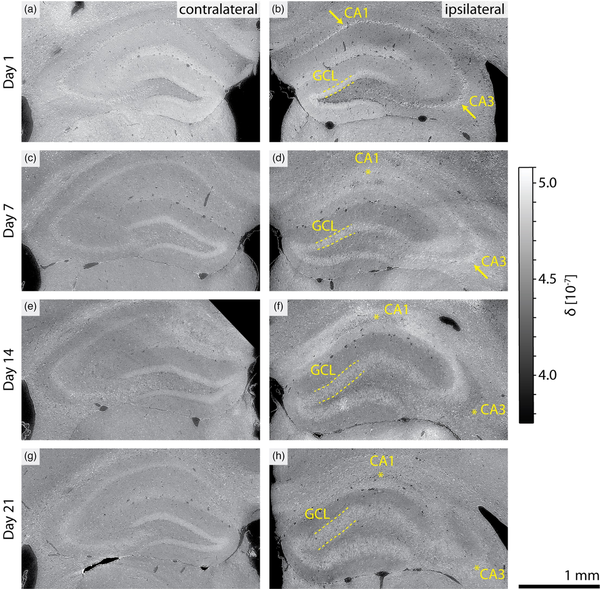
3D X-ray histology reveals histopathological changes in mTLE. Indications of hippocampal sclerosis are observed with increasing time following the KA injection (top to bottom) in virtual coronal slices from selected mice. Neuron loss in the pyramidal layer of the CA1 region (arrow: intact, star: neuronal loss) is observed on Day 7 after the KA injection (d). Neuronal loss in the CA3 field is observed on Day 14 postinjection (f). Granule cell dispersion, characterized by an increased thickness in the granular cell layer (GCL, dashed line), is more prominent for Days 14 and 21 postinjection (f and h). Hippocampal sclerosis is not observed in the contralateral brain hemispheres (a, c, e, and g), which reveal only normal brain anatomical variations across the population. All images are displayed with an index of refraction mapped to intensity values , see grayscale range on the right.
The GCL appeared as a cell-rich band of relatively high density: on average, the GCL had 9% higher intensity than the molecular layer. GCD was identified via an increasing thickness and decreasing mean density of the GCL. The degree of GCD, i.e., GCL thickness, increased over time, following on from the injection. GCL volume in the ipsilateral hemisphere for one Day 7 mouse was already 65% larger relative to the contralateral hemisphere—an effect visible in the virtual slices of the 3D X-ray histology. By Day 21, the GCL of the ipsilateral hemispheres had an average increase in intensity of 19% and a decrease in volume of 180% compared to the contralateral hemispheres.
It should be noted that compared with other time points, increased contrast between the polymorphic and molecular layers of the DG was observed for both hemispheres of the mice sacrificed on Day 1 after KA injection (Fig. 3, top row). For Day 1 KA-injected mice, the mean intensity of the polymorphic layer was 22 and 13% smaller than that of the molecular layer for the ipsi- and contralateral hemispheres, respectively. For all other groups, polymorphic layer intensity was on average 3 and 1% smaller than that of the molecular layer for the ipsi- and contralateral hemispheres, respectively.
Control mice sacrificed on Day 7 after saline injection were also imaged, as illustrated in Figure 4. No pathology was identified in the control mice, with the ipsilateral hemisphere of the saline-injected mouse resembling the contralateral hemisphere of the KA-injected mice. For example, the CA1 and CA3 regions remained intact and no signs of GCD were detected, cf. the Day 7 post-KA injection mouse in Figure 4. The injection site was identified in the 3D datasets, as shown in Figure 2, where increased tissue density was observed but no further histopathological signs were identified. For the Day 7 postsaline-injected mouse shown in Figure 2, the intensity of the injection site was 239% greater than the surrounding tissue.
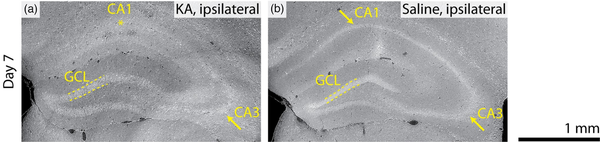

Comparing KA and saline injections. Virtual coronal slices through the 3D X-ray histology of the ipsilateral hemispheres of a mouse sacrificed on Day 7 following KA injection (a) compared with the ipsilateral hemisphere of a mouse sacrificed on Day 7 after saline injection (b). Hippocampal sclerosis is not found in the saline-injected ipsilateral virtual slice (b); however, signs of pyramidal neuron loss in the CA1 region, as well as early signs of granule cell dispersion, are observed for the ipsilateral hemisphere of the KA-injected mouse (a). Note that increased intensity in the Day 7 saline-injected image is related to the injection site (b, below annotation for CA1). Images are displayed with an index of refraction , as in Figure 3.
Benchmarking 3D X-ray Histology Against Histology and Immunohistochemistry
This work represents perhaps the first observation of mTLE pathology with 3D X-ray histology, so validation with gold-standard conventional histology is therefore essential. Immunofluorescent staining subsequent to 3D X-ray histology allowed for the structural confirmation of hippocampal sclerosis, as indicated in Figure 5. NeuN-Iba1 double-immunostained sections confirmed pyramidal neuron loss in the Cornu ammonis and microglia proliferation, indicated with asterisks (*). GCD was confirmed through increasing thickness of the GCL and a diluted NeuN. Within the GCL, electron density (or ) observed in the 3D X-ray histology correlated well with the intensity of the NeuN signal. Supplementary Figure 1 shows further slices, where the specificity of the immunostaining helped identify pyramidal neuron loss in the CA3 region at Day 7 following the injection. This was also observable through increased density (high intensity) in the label-free 3D X-ray histology.
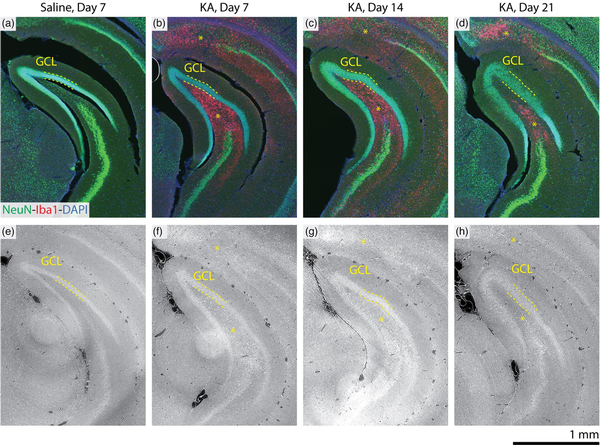
Comparison of 3D X-ray histopathological findings with histology and immunohistochemistry. Optical micrographs of subsequent NeuN-Iba1-DAPI (green-red-blue) immunostained sections (a–d) and virtual slices through the microtomography data (e–h) from a comparable position within the corresponding mouse brain are shown. The fluorescent-labeled sections confirm neuronal loss (diminished green NeuN signal) and indicate microglia activation (increased red Iba1 signal) in the CA1 region, see upper asterisks in b–d, f–h. GCD is indicated by the increased thickness and reduced fluorescence signal (diminished green NeuN signal) of the granular cell layer (GCL, dashed lines). Microglia activation is also confirmed in the polymorph layer of the dentate gyrus. Grayscale for the 3D X-ray histology images is displayed with the index of refraction , as in Figure 3.
The electron-density-based contrast of label-free 3D X-ray histology was benchmarked against common histological and immunofluorescent stains. Figure 6 compares the 3D X-ray histology of a KA-injected mouse 14 days postinjection with photomicrographs of the same brain hemisphere after subsequent sectioning and staining with hematoxylin and eosin (H&E), Nissl, NeuN-Iba1-DAPI, or GFAP-DAPI. In terms of the histochemical stains, hematoxylin stains cell nuclei, eosin stains extracellular matrices and cytoplasms, and Nissl stains cell bodies. For the immunofluorescent stains, NeuN labels neurons, Iba1 labels microglia, GFAP labels astrocytes, and DAPI labels DNA. The density-related contrast of 3D X-ray histology shows correlation with some of the stains; for example, the 3D X-ray histology with inverted look-up table resembles Nissl staining. The increased intensity of the GCL and CA3 layers within the 3D X-ray histology also correlates with NeuN intensity. As histological sections were 50 μm in thickness, an average of 30 consecutive, 1.625 μm-thick slices from the 3D X-ray histology data is also given. This increases the contrast-to-noise ratio of the 3D X-ray histology and makes the appearance more comparable with histology. It should also be noted that strain during sectioning and preparation of histological slides introduced artifacts; for example, a gap opened between the granular and polymorphic layers of the DG as well as between the DG and thalamus.
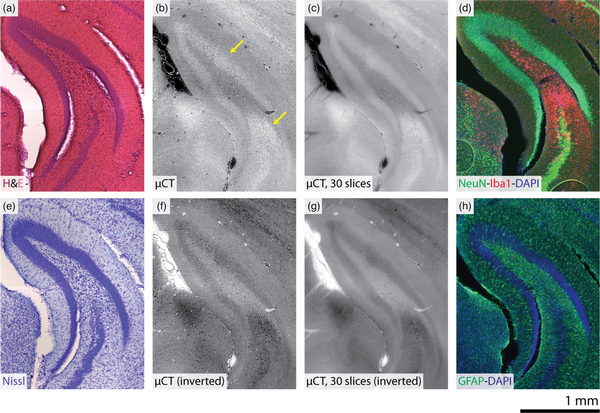
Benchmarking label-free 3D X-ray histology against histological and immunofluorescent stainings. Following X-ray imaging, brain hemispheres were sectioned and sequentially stained with H&E (a), Nissl (e), NeuN-Iba1-DAPI (d), and GFAP-DAPI (f). Optical micrographs of stained sections (50 μm) are compared with virtual coronal sections in a similar position from the 3D X-ray histology from the ipsilateral hemisphere of a mouse on Day 14 after KA injection (b). The z-average of a 49 μm-thick virtual section (μ m slices) (c) and intensity-inverted images are also given for the microtomography dataset (f and g, respectively). Arrows indicate the granular cell layer (upper) and microglial-rich regions (lower), both of which have high intensity in the microtomography dataset (b). Grayscale ranges for the 3D X-ray histology images are displayed with the index of refraction , as in Figure 3.
Convolutional Neural Network-based Segmentation of the Dentate Gyrus
X-ray microtomography generates inherently three-dimensional datasets, thereby providing a unique opportunity to study morphological changes associated with mTLE. Though the anatomy is identifiable in the 3D X-ray histology, fully automatic segmentation of the hippocampal formation, based on the density-related contrast, is impossible. The isotropic, high-resolution nature of 3D X-ray histology presents an additional challenge, as segmenting about 2000 of the 1.625 μm-thick virtual sections per brain hemisphere is prohibitively time-consuming. Here, binned datasets, i.e., (6.5 μm) voxels, were used, as individual cell identification was not needed and binning-related contrast gains (; ) accelerated manual segmentation. The manual segmentation of a limited number of two-dimensional virtual coronal slices was feasible, and in this case the three layers of the DG were labeled in up to 3 min for a single virtual slice. However, with 22 measured brain hemispheres consisting of 514 binned slices each, a semi-automatic approach was highly desirable.
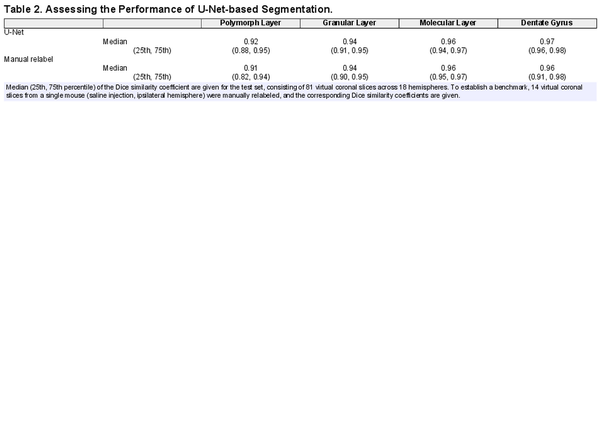
A U-Net-based approach was applied to virtual coronal slices from the 3D X-ray histology, with three-dimensional segmentations produced through slice-by-slice segmentation of stacks, as shown in Figure 7. The accuracy of the two-dimensional U-Net-based segmentation was assessed with the Dice similarity coefficient (DSC) of the test set, consisting of 81 slices from 18 brain hemispheres, as illustrated in Table 2. To determine the gold-standard performance, 14 coronal slices of the ipsilateral hemisphere from a Group 1 mouse were manually relabeled. The DSC with the original manually labeled slices is also given in Table 2. The performance of the U-Net matched that of manual relabeling, and so the two-dimensional performance of the U-Net was deemed satisfactory.
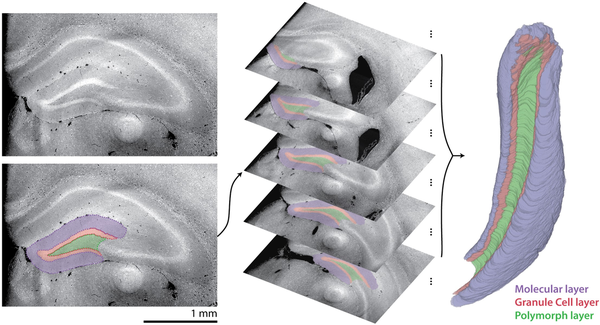
Fig. 7
Segmentation of the dentate gyrus, including molecular, granular, and polymorphic layers. Deep convolutional neural network-based segmentation was performed on virtual coronal sections, here demonstrated on the ipsilateral hemisphere of a mouse 7 days after saline injection (left). Stacks of virtual coronal sections were segmented (center) to produce a three-dimensional segmentation (represented with volume rendering, right).
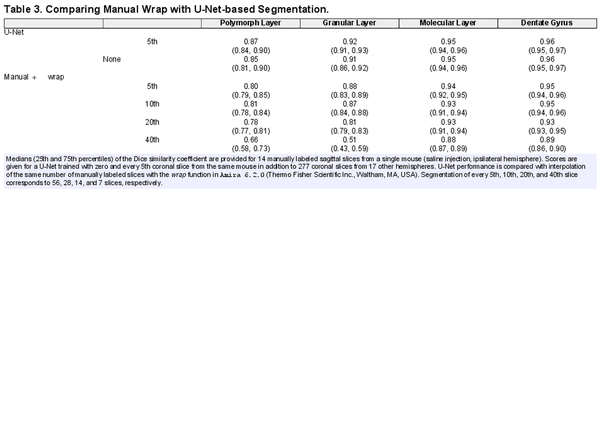
Visual inspection indicated that the resulting volumes from slice-wise segmentation were reasonable—see e.g. Figure 7. To ensure the consistency of the three-dimensional segmentations, accuracy was measured by the DSC with gold-standard segmentations of 14 sagittal slices of the ipsilateral hemisphere from a Group 1 mouse. We benchmarked these results against a three-dimensional interpolation of equally spaced two-dimensional coronal segmentations based on Amira’s wrap function. We additionally compared the effect of training the U-Net with zero and every 5th ground truth virtual coronal slice. Segmentation of every 5th, 10th, 20th, and 40th slice corresponded to 56, 28, 14, and 7 slices, respectively. The results are provided in Table 3. As expected, the manual + wrap approach increased in quality as more slices were labeled, though it provided diminishing returns for segmenting more than every 10th slice. On the other hand, the U-Net approach only slightly improved with more training slices from the brain hemisphere in question, though it should be noted that the U-Net’s training set also contained 277 coronal slices of 17 hemispheres from other mice (median of 20 slices per brain hemisphere). The U-Net outperformed the manual wrap approach in all cases. Therefore, the three-dimensional segmentations produced by two-dimensional U-Net segmentations were considered accurate.
Surprisingly, the U-Net performed well even when no slices from the mouse in question were used in training, cf. Table 3 row “U-Net none.” Thus, at least for the pathology-free Group 1 mice, generalizability of the U-Net was satisfactory. This indicates that the U-Net-based approach scales well to larger sample sizes.
DSC scores give an indication of an overlap between U-Net predictions and gold-standard manual segmentations, but high DSC scores do not necessarily ensure that measured volumes and intensities are equivalent. Therefore, Table 4 provides the median (25th and 75th percentiles) errors as a percentage of the measured volume, mean intensity, and standard deviation of intensity. For each of the 18 samples present in the 81-slice test set, slices were pooled and errors assessed as a percentage of the gold standard, i.e., .
MTLE-induced Volume and Density Changes
Volume changes within the three DG layers post-KA injection were assessed based on the three-dimensional segmentations. Figure 8 shows the volume of the polymorphic, granular, and molecular layers of the DG as well as the total of the three for both ipsilateral and contralateral hemispheres. Note that in accordance with Janz et al. (), positions from to 1820 μm along the rostro-caudal hippocampal axis were considered for volume calculations. This represents the extent of the region where GCD was observed after the KA injection and was consistent in all datasets. The three regions showed increased volumes in the ipsilateral hemisphere, from the time of the KA injection through to sacrifice. As expected from the observed GCD (see Fig. 3), the volume of the granular layer substantially increased as mTLE progressed. It should be noted that the volumes of the polymorphic and molecular layers also increased. The result was a doubling of ipsilateral DG volume for the Day 21 post-KA injection mice compared to both the Day 7 postsaline injection and the Day 1 post-KA injection mice.
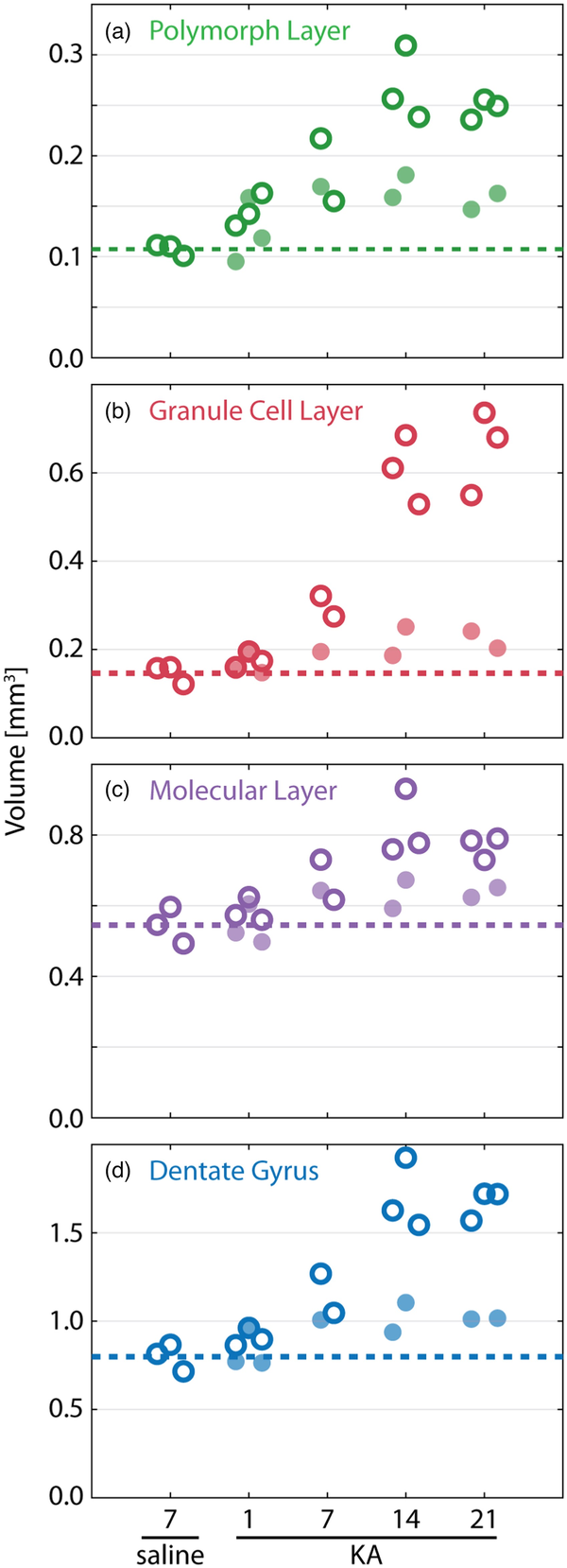
Fig. 8
Volume changes within the dentate gyrus (DG) after KA injection. Volumes of the segmented polymorphic (a), granular cell (b), and molecular layers (c) as well as the full DG (d) are plotted for Day 7 after the saline injection as well as Days 1, 7, 14, and 21 after the KA injection (left to right). Light filled circles (dark rings) correspond to the contralateral (ipsilateral) hemisphere.
The volume difference between the ipsilateral and contralateral hemispheres is shown in Figures 9a–9d for mice where both hemispheres were measured. The results indicate that the ipsilateral volume of the three DG layers increased over time following the injection. The volume of the granular cell layer in the ipsilateral hemisphere doubled with respect to the contralateral hemisphere for mice at Days 14 and 21 postinjection. Image intensity values within the three segmented regions of the DG were also assessed as a proxy for tissue density. Note that the measurements are directly comparable because they were taken at the same beamline with the same measurement parameters, i.e., the same detector, photon statistics, etc. The middle (e–g) and bottom rows (i–l) of Figure 9 plot the difference in the mean and standard deviation of the intensity values between the ipsilateral and contralateral hemispheres for the polymorphic, granular, and molecular layers as well as the full DG (left to right). Mean intensity within the DG layers decreased on the ipsilateral side over time, from injecting the KA through to sacrifice. This is particularly evident within the granular layer, which is consistent with GCD and correlates with the decreased NeuN fluorescent signal intensity observed in the GCL, cf. Figure 5. It should be noted that the mean and standard deviation of the intensity values reveal large variability in the mice sacrificed on Day 1 after the KA injection. Decreasing mean and standard deviations of gray values within the ipsilateral relative to the contralateral hemisphere are observed more clearly in the mice sacrificed on Days 7, 14, and 21 after the KA injection. The standard deviations of the intensity values were elevated on the ipsilateral side on Day 1 after injection, with the effect disappearing 21 days thereafter. The increased standard deviation in the Day 1 post-KA injection mice is likely related to cell death.
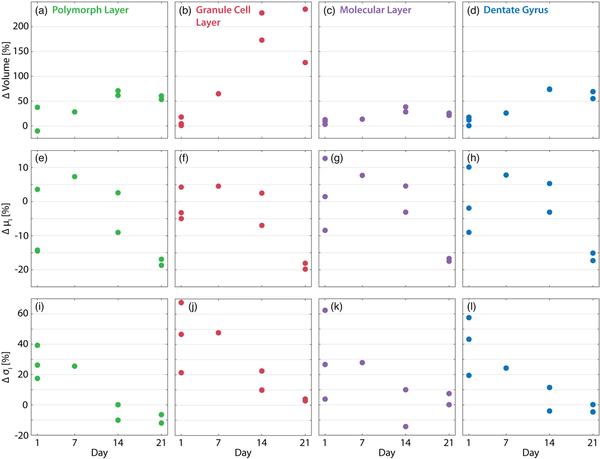
Volume and density changes within the dentate gyrus (DG) after KA injection. The difference in volume (a–d), mean intensity (e–h), and standard deviation of intensity (i–l) between the ipsilateral and contralateral hemispheres are plotted for the segmented polymorphic, granular, and molecular layers as well as the full DG (left to right). Values are given in percentages, i.e., . Note that, in general, inter-animal variability has a substantially larger effect on these metrics than segmentation errors, cf. Table 4.
Discussion
The KA mouse model for mTLE has been extensively validated (; ; ; , ; ), therefore in this study the histopathological changes are surrogate markers for the development of mTLE.
Counting structures such as cells in histological slices has been shown to be prone to a number of errors originating from diverse sources (). Consequently, two-dimensional morphometry e.g. cell counting, based on histological or immunohistochemical slices does not correlate to the more accurate three-dimensional stereological microscopic morphometry, which has been widely developed only in the last 20 years (; ). Depending on the biological or medical question at hand, as few as three-to-five to as many as 30–50 two-dimensional histological slices of a tissue must be prepared and analyzed to reach acceptable precision for the three-dimensional estimates (; ). Combined with multiple histological or immunohistochemical staining required for studies, cf. Janz et al. (), tens or hundreds of histological sections may be needed. This affects not only model animal studies but also leads to important clinical implications and has been a subject of rigorous debate in clinical cases where histopathology plays a pivotal role in diagnosis and therapy planning (). Thus, quantification of anatomical structures of interest down to the cellular and even subcellular level (; ; , ) could be enhanced by the inherent three-dimensional nature of X-ray microtomography. 3D X-ray histology can be performed at an intermediate step of the histological preparation before physical slicing, thus it has been used to guide selection of slicing planes for histological investigation (). X-ray microtomography may also allow for “virtual staining” of three-dimensional volumes with common histological stains ().
3D X-ray histology is an inherently three-dimensional imaging modality, allowing for the quantification of morphological changes in the DG as a result of mTLE. Measurements can be done at any stage of tissue preparation () and without the need for physical sectioning to micrometer thickness including the related slice distortions (). Note that both histology and 3D X-ray histology suffer from deformations due to paraffin embedding, see Rodgers et al. (, ). The volumetric datasets from 3D X-ray histology are advantageous in preclinical studies, as they can be oriented arbitrarily in three-dimensional space e.g. for alignment with brain atlases, visualization of the injection site (see Fig. 2), or for precisely locating electrode positions. Quantifying the degree of hippocampal sclerosis in resected hippocampus from human patients plays an important role in outcome prognosis, but variations along the hippocampal axis complicate any evaluation with limited sections from conventional histology. Thus, volumetric metrics of the degree of hippocampal sclerosis should prove more robust. For future studies, three-dimensional, nonrigid registration should allow for correcting tissue deformations while extracting the skull (), as well as tissue fixation () and embedding (, ). Eventually, correspondence between 3D X-ray histology and the in vivo state could be achieved with multi-modal registration to comparatively lower spatial resolution imaging techniques, including MRI with spatial resolution down to 10 μm resolution (; ; ).
We employed single-distance propagation-based phase contrast with Paganin’s filter (). The resulting contrast is related to electron density through the real part of the index of refraction, as opposed to highly specific histochemical or immunofluorescent stainings that target proteins. In line with previous studies (), immunohistochemistry for different cell types revealed characteristic spatiotemporal patterns during disease progression, including an early neuronal loss in CA1 concomitant to dense microglial scaring and later more widespread reactive astrogliosis and pronounced granule cell dispersion. While X-ray microtomography did not allow detecting cell-type-specific microstructural alterations, histopathological changes such as hippocampal sclerosis could be identified on the basis of the electron density contrast, as illustrated in Figure 3. Electron density correlates with the presence of cells, which is supported by the comparison of the 3D X-ray histology with common stainings, as shown in Figure 6. For example, a decrease in the mean intensity of the GCL, as noted Figure 9 (middle row), can serve as an indication of GCD, which is analogous to a decrease in the NeuN fluorescence signal. Also, an increase in the standard deviation of intensities in the ipsilateral hemisphere on Day 1 postinjection may be linked to cell death. The microanatomical basis for the overall decrease in DG density over time following the injection should be further investigated. This substantial change within the entire DG should be correlated with future magnetic resonance studies. Despite these observable changes, unstained 3D X-ray histology alone was insufficient to identify definitively astrocytes (labeled with GFAP) or differentiate between neurons (NeuN) and microglia (Iba1) (). While the development of X-ray contrast agents may help these shortcomings, label-free imaging of paraffin-embedded tissue offers consistent and easily repeatable results, which is an advantage over sectioning and staining, where variations in section thickness or staining intensity lead to challenges for automated analysis (; ). As demonstrated in this study, more specific conventional histology can be subsequently added to 3D X-ray histology for benchmarking against a wide range of well-established stains.
Slice-to-volume registration between histological sections and volumetric images, which is the process of finding dense spatial correspondences between the two modalities, is a challenging task due to the large search space, limited two-dimensional data, appearance differences due to multi-modality, and topology changes (e.g. cracks) (). Here, histology-to-3D X-ray histology matching may prove simpler than histology-to-MRI matching, thanks to the comparable resolution and more similar appearance between label-free 3D X-ray histology and common stains, cf. Figure 6. In this study, correspondence was only found approximately through the manual rotation and translation of 3D X-ray histology datasets. Furthermore, even after the coarse alignment of 3D X-ray histology with the atlas coordinate frame, several rotations were necessary for approximate alignment with coronally sectioned histology slices. The development of automatic or semi-automatic slice-to-volume registration is therefore desirable, as it would unlock the possibility of virtual staining () of label-free, 3D X-ray histology.
Morphological analysis with sparse, artifact-containing two-dimensional histology slices only allows for approximating the volumes and surface areas of cell layers. These approximations can be refined with 3D X-ray histology, cf. Figure 8 and reference (), Figure 1c. Additionally, measured areas in histological sections e.g. the GCL area for quantifying GCD, depend on the angle of sectioning. Volume measurements may therefore provide more robust outcomes, albeit a detailed comparison is needed with precise correspondence of histological slice and 3D X-ray histology volume, as well as the quantification of segmentation errors for both modalities. In the present study, differences in measured volumes and intensities between ground truth and U-Net-based segmentations were of the order of a few percentage points. For reference, the changes seen in Figure 9 were at least an order of magnitude larger. The three-dimensional nature of 3D X-ray histology should also allow for the introduction of further morphological metrics for quantifying hippocampal sclerosis.
In this study, we calculated the volume of the three DG layers based on segmentation. Simple automated approaches to segmenting unstained 3D X-ray histology data were suboptimal due to a lack of specificity. Anatomical borders are often based on connectivity or cell type in a given region, which were inaccessible in the present 3D X-ray histology data; thus, manual segmentation was needed. It should be kept in mind that even manual segmentation produced variability, as demonstrated by the DSC for manually relabeled slices. In addition, a complete manual segmentation of the 3D X-ray histology data would be extremely time-consuming, so a machine learning-based approach was employed based on a U-Net architecture () and a training set of a limited number of manually segmented two-dimensional virtual slices. This approach produced accurate segmentations with less than a tenth of the volume used for training. Furthermore, it revealed a promising degree of generalizability, as evidenced by Table 3, which is critical for future studies with larger sample sizes. Thus, off-the-shelf networks can accelerate the segmentation of 3D X-ray histology data, and a more tailored network would likely allow for even fewer manually labeled slices, albeit at the expense of development time and computational resources. Based on visual inspection, ring artifacts present in the tomography datasets had only minor effects on segmentation quality, see the examples in Supplementary Figure 2. Note that also the manually labeled training data included slices with such artifacts. Due to their relative infrequency within the volumetric data and independence from dentate gyrus region, hemisphere, and group, ring artifacts are not expected to introduce bias but may somewhat increase variance of measured quantities. Thus, errors in the mean or standard deviation of intensity values are expected to be smaller than those due to segmentation errors reported in Table 4. Volume errors due to ring artifacts are encompassed in the values of Table 4 because manual gold-standard labels did not suffer from segmentation errors in these regions.
In the present study, both the morphology and density of the DG layers showed changes during epileptogenesis. Future studies with higher resolution will uncover subcellular features, which will not only help improve segmentation (; ) but also allow for a more detailed understanding of the pathological progression. Segmentation of individual cells (; ; ) will allow for testing the hypothesis that an increased standard deviation of the signal intensity from Day 1 mice is related to cell death.
A drawback of the present synchrotron radiation-based approach is the lack of accessibility. Due to limitations in beamtime, both ipsi- and contralateral hemispheres could be measured for only eight of the 14 mice. Laboratory-based microtomography may bridge this gap, with impressive results using conventional absorption contrast () and more recently phase contrast (, ). As more of these systems enter the market (), 3D X-ray histology can be integrated into the existing workflow of histopathology (). A major practical barrier for these laboratory systems in a clinical setting is long scan times e.g. more than 10 h for true micrometer resolution. Future studies should examine how far the increased flux from advanced X-ray sources, such as liquid metal jet systems, can address this issue.
For future mTLE studies, three-dimensional information from 3D X-ray histology will allow for assessing the spatial extent of tissue damage and scarring at the kainate injection site, thus guiding a limited number of histological sections in the most relevant regions. Further, precise structural information is crucial for interpretation of electrophysiological signals. Thus, microanatomical information from 3D X-ray histology could aid EEG studies with planning electrode positioning or precise retrospective determination of electrode locations.
MRI plays an essential role in investigations of mTLE, particularly since it is the gold standard for in vivo imaging. It provides structural and functional information through techniques like functional MRI or diffusion tensor imaging, while also offering the potential for multi-parametric imaging to combine anatomical, functional, and metabolic information in a single scanning session. While variations of MRI can achieve spatial resolutions on the order of tens of micrometers, post mortem X-ray microtomography with true micrometer resolution provides higher resolution by an order of magnitude higher in all three spatial dimensions, i.e., voxels more than 10 times smaller. 3D X-ray histology provides structural imaging of length scales from cellular-level, including neuronal morphology, for macroscopic brain regions or even entire brains (). 3D X-ray histology is therefore uniquely positioned to bridge the gap between nondestructive, including in vivo three-dimensional MRI and two-dimensional histological analysis at the cellular level (). Future studies should include a quantitative comparison of MRI and 3D X-ray histology, focusing on whether the higher resolution of the latter improves quantification of three-dimensional structures. Notably, although previous high-resolution MRI in the intrahippocampal mouse model for mTLE successfully tracked the progression of histological changes primarily associated with granule cell dispersion (), it lacked clear cellular resolution. However, the ability to deduce cell densities in hippocampal subfields holds great significance for mTLE research, considering the distinct functions of cell subtypes, such as pyramidal cells in the Cornu ammonis, dentate granule cells, and hilar mossy cells. This cellular information could assist researchers in unraveling how the loss of neurons in these specific subfields contributes to the occurrence of epileptiform discharges and their distinctive characteristics.
For treatment of mTLE in a clinical setting, MRI plays a crucial role in epilepsy surgery planning, while histology is vital for postoperative analysis of hippocampal sclerosis to assess the long-term prognosis of seizure freedom. Interestingly, microstructural features that may be inaccessible with MRI appear to yield predictive value for postsurgical outcome (; ; ; ). Still, variations along the hippocampal axis hinder the quantification of the degree of hippocampal sclerosis and complicate any evaluation with limited sections from conventional histology.
Conclusions
We have demonstrated the utility of post mortem 3D X-ray histology for studying epileptogenesis in a mouse model. Histopathological progression was identified via the electron density-based contrast of label-free X-ray microtomography, and three-dimensional morphology was assessed without physical sectioning. Importantly, 3D X-ray histology was benchmarked against gold-standard histopathological analysis based on optical microscopy. Analysis of the volumetric data requires distinct tools compared to conventional histology, though it is possible to take advantage of developments from medical imaging and other microscopy disciplines e.g. three-dimensional electron microscopy. For example, the use of deep convolutional neural networks has enabled microstructural analysis without exhaustive manual labeling. Thus, 3D X-ray histology complements conventional histopathological analysis and unlocks the third dimension. This X-ray microscopy technique will find increasing application in epilepsy research and beyond.
Availability of Data
3D X-ray histology data for all mice are available in an EBRAINS repository (doi.org/10.25493/3J7E-E0P). Downsampled data, gold-standard manual segmentations, and trained network parameters are also available at this link. The source code for the segmentation neural network is available from https://github.com/grodgers1/EpilepsySegmentation.
Supplementary Material
To view supplementary material for this article, please visit https://doi.org/10.1093/micmic/ozad082.
Acknowledgments
Beamtime for this experiment was provided by the European Synchrotron Radiation Facility (ESRF) through application MD 896. The authors thank A. Rack and the staff at beamline ID19 of ESRF for their support during the beamtime.
References
- Arabadzisz D, Antal K, Parpan F, Emri Z, Fritschy J-M (2005). Epileptogenesis and chronic seizures in a mouse model of temporal lobe epilepsy are associated with distinct EEG patterns and selective neurochemical alterations in the contralateral hippocampus. Exp Neurol194(1), 76–90. https://doi.org/10.1016/j.expneurol.2005.01.029
- Balwani A, Miano J, Liu R, Kitchell L, Prasad JA, Johnson EC, Gray-Roncal W, Dyer EL. (2021). Multi-scale modeling of neural structure in X-ray imagery. In 2021 IEEE International Conference on Image Processing (ICIP), pp. 141–145. Available at https://doi.org/10.1109/ICIP42928.2021.9506174.
- Bancroft JD, Gamble M (2008). Theory and Practice of Histological Techniques, 6th ed. Philadelphia, PA: Churchill Livingstone Elsevier.
- Bratu VA, Erpenbeck VJ, Fehrenbach A, Rausch T, Rittinghausen S, Krug N, Hohlfeld JM, Fehrenbach H (2014). Cell counting in human endobronchial biopsies: Disagreement of 2D versus 3D morphometry. PLoS One9(3), e92510. https://doi.org/10.1371/journal.pone.0092510
- Carroll ML, Carroll NG, James AL (2006). Do bronchial biopsies represent mast cell density in airways? A stereological study. Eur Respir J28(3), 612–621. https://doi.org/10.1183/09031936.06.00037006
- Danka T. (2020). pytorch-unet. Available at https://github.com/cosmic-cortex/pytorch-UNet.
- Deleo F, Garbelli R, Milesi G, Gozzo F, Bramerio M, Villani F, Cardinale F, Tringali G, Spreafico R, Tassi L (2016). Short-and long-term surgical outcomes of temporal lobe epilepsy associated with hippocampal sclerosis: Relationships with neuropathology. Epilepsia57(2), 306–315. https://doi.org/10.1111/epi.13277
- Engel J (2001). Mesial temporal lobe epilepsy: What have we learned?Neuroscientist7(4), 340–352. https://doi.org/10.1177/107385840100700410
- Engel J (2006). Report of the ILAE classification core group. Epilepsia47(9), 1558–1568. https://doi.org/10.1111/j.1528-1167.2006.00215.x
- Farhoodi R, Lansdell BJ, Kording KP (2019). Quantifying how staining methods bias measurements of neuron morphologies. Front Neuroinform13, 36. https://doi.org/10.3389/fninf.2019.00036
- Ferrante E, Paragios N (2017). Slice-to-volume medical image registration: A survey. Med Image Anal39, 101–123. https://doi.org/10.1016/j.media.2017.04.010
- Flint JJ, Lee CH, Hansen B, Fey M, Schmidig D, Bui JD, King MA, Vestergaard-Poulsen P, Blackband SJ (2009). Magnetic resonance microscopy of mammalian neurons. NeuroImage46(4), 1037–1040. https://doi.org/10.1016/j.neuroimage.2009.03.009
- Grote A, Heiland D-H, Taube J, Helmstaedter C, Ravi VM, Will P, Hattingen E, Schüre J-R, Witt J-A, Reimers A, Elger C, Schramm J, Becker AJ, Delev D (2023). ‘Hippocampal innate inflammatory gliosis only’ in pharmacoresistant temporal lobe epilepsy. Brain146(2), 549–560. https://doi.org/10.1093/brain/awac293
- Guillery RW (2002). On counting and counting errors. J Comp Neurol447(1), 1–7. https://doi.org/10.1002/cne.10221
- Häussler U, Bielefeld L, Froriep UP, Wolfart J, Haas CA (2012). Septotemporal position in the hippocampal formation determines epileptic and neurogenic activity in temporal lobe epilepsy. Cereb Cortex22(1), 26–36. https://doi.org/10.1093/cercor/bhr054
- Heinrich C, Nitta N, Flubacher A, Müller M, Fahrner A, Kirsch M, Freiman T, Suzuki F, Depaulis A, Frotscher M, Haas CA (2006). Reelin deficiency and displacement of mature neurons, but not neurogenesis, underlie the formation of granule cell dispersion in the epileptic hippocampus. J Neurosci26(17), 4701–4713. https://doi.org/10.1523/JNEUROSCI.5516-05.2006
- Hieber SE, Bikis C, Khimchenko A, Schweighauser G, Hench J, Chicherova N, Schulz G, Müller B (2016). Tomographic brain imaging with nucleolar detail and automatic cell counting. Sci Rep6(1), 32156. https://doi.org/10.1038/srep32156
- Hsia CC, Hyde DM, Ochs M, Weibel ER (2010). An official research policy statement of the American Thoracic Society/European Respiratory Society: Standards for quantitative assessment of lung structure. Am J Respir Crit Care Med181(4), 394–418. https://doi.org/10.1164/rccm.200809-1522ST
- Janz P, Hauser P, Heining K, Nestel S, Kirsch M, Egert U, Haas CA (2018). Position- and time-dependent arc expression links neuronal activity to synaptic plasticity during epileptogenesis. Front Cell Neurosci12, 244. https://doi.org/10.3389/fncel.2018.00244
- Janz P, Savanthrapadian S, Häussler U, Kilias A, Nestel S, Kretz O, Kirsch M, Bartos M, Egert U, Haas CA (2017a). Synaptic remodeling of entorhinal input contributes to an aberrant hippocampal network in temporal lobe epilepsy. Cereb Cortex27(3), 2348–2364. https://doi.org/10.1093/cercor/bhw093
- Janz P, Schwaderlapp N, Heining K, Häussler U, Korvink JG, von Elverfeldt D, Hennig J, Egert U, LeVan P, Haas CA (2017b). Early tissue damage and microstructural reorganization predict disease severity in experimental epilepsy. eLife6, e25742. https://doi.org/10.7554/eLife.25742
- Johnson AM, Sugo E, Barreto D, Hiew C-C, Lawson JA, Connolly AM, Somerville E, Hasic E, Bye AM, Cunningham AM (2016). The severity of gliosis in hippocampal sclerosis correlates with pre-operative seizure burden and outcome after temporal lobectomy. Mol Neurobiol53, 5446–5456. https://doi.org/10.1007/s12035-015-9465-y
- Kak AC, Slaney M (Principles of Computerized Tomographic Imaging. Classics in Applied Mathematics. Society for Industrial and Applied Mathematics. 2001). Available at https://doi.org/10.1137/1.9780898719277.
- Khimchenko A, Bikis C, Pacureanu A, Hieber SE, Thalmann P, Deyhle H, Schweighauser G, Hench J, Frank S, Müller-Gerbl M, Schulz G, Cloetens P, Müller B (2018). Hard X-ray nanoholotomography: Large-scale, label-free, 3D neuroimaging beyond optical limit. Advanced Science5(6), 1700694. https://doi.org/10.1002/advs.201700694
- Khimchenko A, Deyhle H, Schulz G, Schweighauser G, Hench J, Chicherova N, Bikis C, Hieber SE, Müller B (2016). Extending two-dimensional histology into the third dimension through conventional micro computed tomography. NeuroImage139, 26–36. https://doi.org/10.1016/j.neuroimage.2016.06.005
- Kingma DP, Ba J (2017). Adam: A method for stochastic optimization. arXiv, arXiv:1412.6980, preprint: not peer reviewed.
- Klein S, Staring M, Murphy K, Viergever MA, Pluim JPW (2010). Elastix: A toolbox for intensity-based medical image registration. IEEE Trans Med Imaging29(1), 196–205. https://doi.org/10.1109/TMI.2009.2035616
- Kowalski J, Geuting M, Paul S, Dieni S, Laurens J, Zhao S, Drakew A, Haas CA, Frotscher M, Vida I (2010). Proper layering is important for precisely timed activation of hippocampal mossy cells. Cereb Cortex20(9), 2043–2054. https://doi.org/10.1093/cercor/bhp267
- Kuan AT, Phelps JS, Thomas LA, Nguyen TM, Han J, Chen C-L, Azevedo AW, Tuthill JC, Funke J, Cloetens P, Pacureanu A, Lee W-CA (2020). Dense neuronal reconstruction through X-ray holographic nano-tomography. Nat Neurosci23(12), 1637–1643. https://doi.org/10.1038/s41593-020-0704-9
- Kuo W, Le NA, Spingler B, Wenger RH, Kipar A, Hetzel U, Schulz G, Müller B, Kurtcuoglu V (2020). Simultaneous three-dimensional vascular and tubular imaging of whole mouse kidneys with X-ray μCT. Microsc Microanal26(4), 731–740. https://doi.org/10.1017/S1431927620001725
- LaGrow TJ, Moore MG, Prasad JA, Davenport MA, Dyer EL ( 2018). Approximating cellular densities from high-resolution neuroanatomical imaging data. In 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, pp. 1–4. IEEE. Available at https://doi.org/10.1109/EMBC.2018.8512220.
- Lee CH, Blackband SJ, Fernandez-Funez P (2015). Visualization of synaptic domains in the Drosophila brain by magnetic resonance microscopy at 10 micron isotropic resolution. Sci Rep5(1), 8920. https://doi.org/10.1038/srep08920
- Lein ES, Hawrylycz MJ, Ao N, Ayres M, Bensinger A, Bernard A, Boe AF, Boguski MS, Brockway KS, Byrnes EJ, Chen L, Chen L, Chen T-M, Chi Chin M, Chong J, Crook BE, Czaplinska A, Dang CN, Datta S, Dee NR, Desaki AL, Desta T, Diep E, Dolbeare TA, Donelan MJ, Dong H-W, Dougherty JG, Duncan BJ, Ebbert AJ, Eichele G, Estin LK, Faber C, Facer BA, Fields R, Fischer SR, Fliss TP, Frensley C, Gates SN, Glattfelder KJ, Halverson KR, Hart MR, Hohmann JG, Howell MP, Jeung DP, Johnson RA, Karr PT, Kawal R, Kidney JM, Knapik RH, Kuan CL, Lake JH, Laramee AR, Larsen KD, Lau C, Lemon TA, Liang AJ, Liu Y, Luong LT, Michaels J, Morgan JJ, Morgan RJ, Mortrud MT, Mosqueda NF, Ng LL, Ng R, Orta GJ, Overly CC, Pak TH, Parry SE, Pathak SD, Pearson OC, Puchalski RB, Riley ZL, Rockett HR, Rowland SA, Royall JJ, Ruiz MJ, Sarno NR, Schaffnit K, Shapovalova NV, Sivisay T, Slaughterbeck CR, Smith SC, Smith KA, Smith BI, Sodt AJ, Stewart NN, Stumpf K-R, Sunkin SM, Sutram M, Tam A, Teemer CD, Thaller C, Thompson CL, Varnam LR, Visel A, Whitlock RM, Wohnoutka PE, Wolkey CK, Wong VY, Wood M, Yaylaoglu MB, Young RC, Youngstrom BL, Feng Yuan X, Zhang B, Zwingman TA, Jones AR (2007). Genome-wide atlas of gene expression in the adult mouse brain. Nature445(7124), 168–176. https://doi.org/10.1038/nature05453
- Lévesque M, Avoli M (2013). The kainic acid model of temporal lobe epilepsy. Neurosci Biobehav Rev37(10), 2887–2899. https://doi.org/10.1016/j.neubiorev.2013.10.011
- Marx M, Haas C, Häussler U (2013). Differential vulnerability of interneurons in the epileptic hippocampus. Front Cell Neurosci7, 167. https://doi.org/10.3389/fncel.2013.00167
- Meldrum BS, Vigouroux RA, Rage P, Brierley JB (1973). Hippocampal lesions produced by prolonged seizures in paralyzed artificially ventilated baboons. Experientia29(5), 561–563. https://doi.org/10.1007/BF01926665
- Migga A, Schulz G, Rodgers G, Osterwalder M, Tanner C, Blank H, Jerjen I, Salmon P, Twengström W, Scheel M, Weitkamp T, Schlepütz CM, Bolten JS, Huwyler J, Hotz G, Madduri S, Müller B (2022). Comparative hard X-ray tomography for virtual histology of zebrafish larva, human tooth cementum, and porcine nerve. J Med Imaging9(3), 031507. https://doi.org/10.1117/1.JMI.9.3.031507
- Milligan K, Balwani A, Dyer E (2019). Brain mapping at high resolutions: Challenges and opportunities. Curr Opin Biomed Eng12, 126–131. https://doi.org/10.1016/j.cobme.2019.10.009
- Paganin D, Mayo SC, Gureyev TE, Miller PR, Wilkins SW (2002). Simultaneous phase and amplitude extraction from a single defocused image of a homogeneous object. J Microsc206(1), 33–40. https://doi.org/10.1046/j.1365-2818.2002.01010.x
- Pichat J, Iglesias JE, Yousry T, Ourselin S, Modat M (2018). A survey of methods for 3D histology reconstruction. Med Image Anal46, 73–105. https://doi.org/10.1016/j.media.2018.02.004
- Prasad JA, Balwani AH, Johnson EC, Miano JD, Sampathkumar V, De Andrade V, Fezzaa K, Du M, Vescovi R, Jacobsen C, Kording KP, Gürsoy D, Gray Roncal W, Kasthuri N, Dyer EL (2020). A three-dimensional thalamocortical dataset for characterizing brain heterogeneity. Sci Data7(1), 358. https://doi.org/10.1038/s41597-020-00692-y
- Rau C, Weitkamp T, Snigirev A, Schroer CG, Tümmler J, Lengeler B (2001). Recent developments in hard X-ray tomography. Nucl Instrum Methods Phys Res A467–468, 929–931. https://doi.org/10.1016/S0168-9002(01)00528-9
- Riban V, Bouilleret V, Pham-Lê BT, Fritschy JM, Marescaux C, Depaulis A (2002). Evolution of hippocampal epileptic activity during the development of hippocampal sclerosis in a mouse model of temporal lobe epilepsy. Neuroscience112(1), 101–111. https://doi.org/10.1016/S0306-4522(02)00064-7
- Rivenson Y, Wang H, Wei Z, de Haan K, Zhang Y, Wu Y, Günaydın H, Zuckerman JE, Chong T, Sisk AE, Westbrook LM, Wallace WD, Ozcan A (2019). Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat Biomed Eng3(6), 466–477. https://doi.org/10.1038/s41551-019-0362-y
- Rodgers G, Kuo W, Schulz G, Scheel M, Migga A, Bikis C, Tanner C, Kurtcuoglu V, Weitkamp T, Müller B (2021). Virtual histology of an entire mouse brain from formalin fixation to paraffin embedding. Part 1: Data acquisition, anatomical feature segmentation, tracking global volume and density changes. J Neurosci Methods364, 109354. https://doi.org/10.1016/j.jneumeth.2021.109354
- Rodgers G, Schulz G, Deyhle H, Kuo W, Rau C, Weitkamp T, Müller B (2020). Optimizing contrast and spatial resolution in hard X-ray tomography of medically relevant tissues. Appl Phys Lett116(2), 023702. https://doi.org/10.1063/1.5133742
- Rodgers G, Tanner C, Schulz G, Migga A, Kuo W, Bikis C, Scheel M, Kurtcuoglu V, Weitkamp T, Müller B (2022a). Virtual histology of an entire mouse brain from formalin fixation to paraffin embedding. Part 2: Volumetric strain fields and local contrast changes. J Neurosci Methods365, 109385. https://doi.org/10.1016/j.jneumeth.2021.109385
- Rodgers G, Tanner C, Schulz G, Weitkamp T, Scheel M, Alarcón MG, Kurtcuoglu V, Müller B (2022b). Mosaic microtomography of a full mouse brain with sub-micron pixel size. In Developments in X-Ray Tomography XIV, vol. 12242, pp. 323–335. SPIE. Available at https://doi.org/10.1117/12.2633556.
- Ronneberger O, Fischer P, Brox T (2015). U-Net: Convolutional networks for biomedical image segmentation. arXiv, arXiv:1505.04597, preprint: not peer reviewed.
- Rusina E, Bernard C, Williamson A (2021). The kainic acid models of temporal lobe epilepsy. eNeuro8(2), ENEURO.0337-20.2021. https://doi.org/10.1523/ENEURO.0337-20.2021
- Ryvlin P, Kahane P (2005). The hidden causes of surgery-resistant temporal lobe epilepsy: Extratemporal or temporal plus? Editorial review. Curr Opin Neurol18(2), 125–127. https://doi.org/10.1097/01.wco.0000162852.22026.6f
- Sala-Padro J, Miró J, Rodriguez-Fornells A, Quintana M, Vidal N, Plans G, Santurino M, Falip M, Camara E (2020). Hippocampal microstructural architecture and surgical outcome: Hippocampal diffusivity could predict seizure relapse. Seizure76, 84–88. https://doi.org/10.1016/j.seizure.2020.01.006
- Schulz G, Crooijmans HJA, Germann M, Scheffler K, Müller-Gerbl M, Müller B (2011). Three-dimensional strain fields in human brain resulting from formalin fixation. J Neurosci Methods202(1), 17–27. https://doi.org/10.1016/j.jneumeth.2011.08.031
- Schulz G, Waschkies C, Pfeiffer F, Zanette I, Weitkamp T, David C, Müller B (2012). Multimodal imaging of human cerebellum—Merging X-ray phase microtomography, magnetic resonance microscopy and histology. Sci Rep2(1), 1–7. https://doi.org/10.1038/srep00826
- Shalaby AM, Sharaf Eldin HE, Abdelsameea AA, Abdelnour HM, Alabiad MA, Elkholy MR, Aboregela AM (2022). Betahistine attenuates seizures, neurodegeneration, apoptosis, and gliosis in the cerebral cortex and hippocampus in a mouse model of epilepsy: A histological, immunohistochemical, and biochemical study. Microsc Microanal28(5), 1734–1748. https://doi.org/10.1017/S1431927622012107
- Shamonin D, Bron E, Lelieveldt B, Smits M, Klein S, Staring M (2014). Fast parallel image registration on CPU and GPU for diagnostic classification of Alzheimer’s disease. Front Neuroinform7, 50. https://doi.org/10.3389/fninf.2013.00050
- Sierra A, Gröhn O, Pitkänen A (2015). Imaging microstructural damage and plasticity in the hippocampus during epileptogenesis. Neuroscience309, 162–172. https://doi.org/10.1016/j.neuroscience.2015.04.054
- Stalder AK, Ilgenstein B, Chicherova N, Deyhle H, Beckmann F, Mueller B, Hieber SE (2014). Combined use of micro computed tomography and histology to evaluate the regenerative capacity of bone grafting materials. Int J Mater Res105(7), 679–691. https://doi.org/10.3139/146.111050
- Sullivan P, Stephens D, Ansari T, Costello J, Jeffery P (1998). Variation in the measurements of basement membrane thickness and inflammatory cell number in bronchial biopsies. Eur Respir J12(4), 811–815. https://doi.org/10.1183/09031936.98.12040811
- Thalmann P, Bikis C, Schulz G, Paleo P, Mirone A, Rack A, Siegrist S, Cörek E, Huwyler J, Müller B. (2017). Removing ring artefacts from synchrotron radiation-based hard X-ray tomography data. In Developments in X-Ray Tomography XI, vol. 10391, pp. 1039114. International Society for Optics and Photonics. Available at https://doi.org/10.1117/12.2274236.
- Thom M (2014). Review: Hippocampal sclerosis in epilepsy: A neuropathology review. Neuropathol Appl Neurobiol40(5), 520–543. https://doi.org/10.1111/nan.12150
- Thurner P, Beckmann F, Müller B (2004). An optimization procedure for spatial and density resolution in hard X-ray micro-computed tomography. Nuclear Instruments and Methods in Physics Research Section B: Beam Interactions with Materials and Atoms225(4), 599–603. https://doi.org/10.1016/j.nimb.2004.05.027
- Töpperwien M, Markus A, Alves F, Salditt T (2019). Contrast enhancement for visualizing neuronal cytoarchitecture by propagation-based X-ray phase-contrast tomography. NeuroImage199, 70–80. https://doi.org/10.1016/j.neuroimage.2019.05.043
- Töpperwien M, van der Meer F, Stadelmann C, Salditt T (2018). Three-dimensional virtual histology of human cerebellum by X-ray phase-contrast tomography. Proc Natl Acad Sci USA115(27), 6940–6945. https://doi.org/10.1073/pnas.1801678115
- Töpperwien M, van der Meer F, Stadelmann C, Salditt T (2020). Correlative X-ray phase-contrast tomography and histology of human brain tissue affected by Alzheimer’s disease. NeuroImage210, 116523. https://doi.org/10.1016/j.neuroimage.2020.116523
- Twengström W, oro CF, omell J, arsson JC, parrelid E, Björnstedt M, Hertz HM (2022). Can laboratory X-ray virtual histology provide intraoperative 3D tumor resection margin assessment?J Med Imaging9(3), 031503. https://doi.org/10.1117/1.JMI.9.3.031503
- Vasquez SX., Small D., Weinstein D. (2013). Beyond bone: Micro-computed tomography for preclinical specimen imaging and applications for soft tissue discovery and translational therapeutic testing. Microsc Microanal19(S2), 620–621. https://doi.org/10.1017/S1431927613005096
- von Bohlen und Halbach O, Lotze M, Pfannmöller JP (2014). Post-mortem magnetic resonance microscopy (MRM) of the murine brain at 7 Tesla results in a gain of resolution as compared to in vivo MRM. Front Neuroanat8, 47.
- Wagner R, Van Loo D, Hossler F, Czymmek K, Pauwels E, Van Hoorebeke L (2011). High-resolution imaging of kidney vascular corrosion casts with nano-CT. Microsc Microanal17(2), 215–219. https://doi.org/10.1017/S1431927610094201
- Walker MC (2015). Hippocampal sclerosis: Causes and prevention. Semin Neurol35(03), 193–200. https://doi.org/10.1055/s-0035-1552618
- Wang Q, Ding S-L, Li Y, Royall J, Feng D, Lesnar P, Graddis N, Naeemi M, Facer B, Ho A, Dolbeare T, Blanchard B, Dee N, Wakeman W, Hirokawa KE, Szafer A, Sunkin SM, Oh SW, Bernard A, Phillips JW, Hawrylycz M, Koch C, Zeng H, Harris JA, Ng L (2020). The Allen mouse brain common coordinate framework: A 3D reference atlas. Cell181(4), 936–953.e20. https://doi.org/10.1016/j.cell.2020.04.007
- Yushkevich PA, Piven J, Hazlett HC, Smith RG, Ho S, Gee JC, Gerig G (2006). User-guided 3D active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. NeuroImage31(3), 1116–1128. https://doi.org/10.1016/j.neuroimage.2006.01.015